Mutational analysis: on the road to refined standards
LCMC II
The Lung Cancer Mutation Consortium (LCMC) is a multi-institutional consortium for the study of driver mutations of lung adenocarcinoma. The cooperating sites enable the identification of relatively large numbers of patients with uncommon and rare alterations, facilitate the analysis of their clinical characteristics, and lay the ground for targeted therapy trials.
LCMC II, which is the second round for LCMC, started in 2012 [1]. The first round, LCMC I, was initiated in 2009 and demonstrated that genomic profiling can work as a multi-institutional effort for patient benefit. Sixteen sites participated in LCMC II, and 14 selected gene alterations were analysed. By the end of the project, all of the sites had accomplished the transition to next-generation sequencing for mutation identification, which was one of the goals of LCMC II.
The genes studied in LCMC II included point mutations in AKT1, BRAF, EGFR, HER2, KRAS, MAP2K1 (MEK), PIK3CA and NRAS, as well as rearrangements in ALK, RET and ROS1, and other alterations, including METamp, PTENexp and METexp. Patients with stage-IV adenocarcinoma of the lung participated in the project. The results of the gene analyses were reported to the LCMC Virtual Database, as well as to the treating physicians, who could use them to select therapies, either as standard-of-care, or to recommend clinical, agent-specific trials, or off-label therapies. Subsequently, the patient outcomes obtained were reported back to the LCMC Database. Genotyping was performed in 875 patients; 242 of these showed targetable driver alterations. Finally, 131 patients went on to targeted therapy.
Improved survival due to expanded genomic analysis
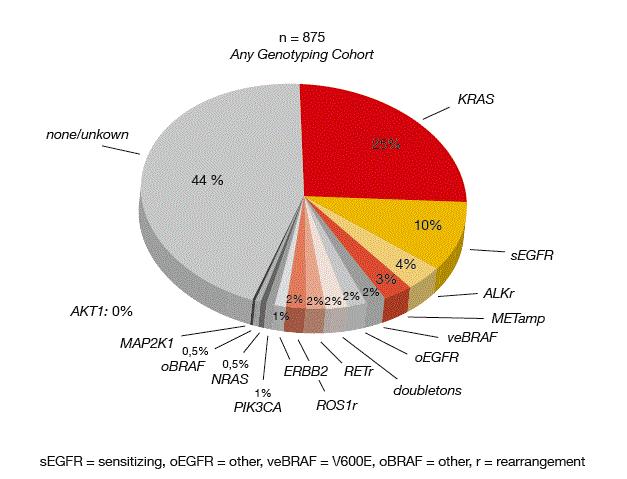
The Figure shows the distribution of the mutations as defined in LCMC II. PTEN loss and MET expression are not included; for these, 15 % and 59 % of cases were positive, respectively. As previously described, overlapping alterations occurred infrequently (4.1 % of total cases).
Figure: Mutation frequencies observed in LCMC II in 875 patients with adenocarcinoma of the lung
Many of the known associations with patient outcomes, such as benefits due to EGFR TKI therapy in the EGFR-mutated population, were observed in LCMC II. Also, smoking status and specific gene alterations correlated with each other, as expected. When all driver mutations were considered, targeted therapy gave rise to survival improvement. On the whole, concomitant mutations in TP53 and/or PTEN and/or PIK3CA did not influence the effects of targeted therapy. Some modulators were identified, however. In the group of EGFR-positive patients treated with TKI therapy, the presence of TP53 had modulatory effects on the benefit of the targeted therapy. Survival probability was higher if the mutation was absent. KRAS mutations were demonstrated to confer worse prognosis in never-smokers.
The authors concluded that expanded molecular testing and associated targeted therapy provides survival benefit, but the assay systems represent an important aspect. Mutation rates obtained with different testing systems can vary widely. For example, TP53 mutation status is most likely under-observed, which limits the ability to detect the affected patients and to develop treatment options. As testing and therapies co-evolve, additional improvements can be expected.
Circulating tumour cells mirror reality
Next-generation sequencing (NGS) of circulating tumour DNA (ctDNA) enables non-invasive profiling of solid tumours. To date, liquid biopsy studies have been limited to modest-sized cohorts and case studies. A large-scale genomic analysis has now established that patterns of genetic changes detected via liquid biopsies can closely mirror changes identified via traditional tumour biopsies [2]. Blood samples were obtained from more than 15,000 patients with advanced-stage cancer of 50 different types. Thirty-seven percent of patients had lung cancer. Somatic genomic profiling was performed using a highly accurate, deep-coverage ctDNA NGS test targeting 70 genes. This is one of the largest cancer genomics studies ever conducted.
With the exception of resistance mutations such as EGFR T790M, the cancer-type-specific frequencies and mutual exclusivity patterns among major driver alterations (as assessed by ctDNA) largely resembled the tissue alteration patterns. When ctDNA was positive for key abnormalities in EGFR, BRAF, KRAS, ALK, RET and ROS1, the same mutations were reported in tissue in 94 % to 100 %. Most ctDNA alterations were detected at very low levels. Half of these occurred at a frequency below 0.4 % of the total DNA in circulation. Even at those low levels, the accuracy of the liquid biopsy remained high. Overall, ctDNA testing revealed a potential targeted treatment option for almost two thirds of the patients tested.
In the NSCLC subset, 51 cases of driver aberrations were detected using ctDNA NGS, in addition to those detected by tissue genotyping. The actionable biomarker yield was thus increased by 42 %. Overall, this analysis illustrated the ability of plasma mutation detection to enhance or complement tissue analysis.
ctDNA as a prognostic marker
Lin et al. hypothesised that tumour-specific alterations in ctDNA quantify tumour heterogeneity and can serve as a non-invasive means to determine prognosis and recurrence in patients with unresectable stage III NSCLC who are treated with curative-intent chemoradiotherapy [3]. Tumour heterogeneity is correlated with therapeutic resistance and poor prognosis. The investigators assessed ctDNA in 156 patients with unresectable NSCLC who were receiving definitive radiotherapy (XRT) or chemo-XRT. Blood was taken before, during, and after therapy. An NGS assay was used to detect single nucleotide variants in 70 genes, amplifications in 16 genes, as well as select fusions and indels.
According to the interim analysis, four main patterns of ctDNA changes were found across serial time-points: specific alterations persistent throughout XRT (n = 9), no alterations in the post-XRT sample (n = 14), increased levels from baseline (n = 10), and alterations that fluctuated throughout therapy (n = 11). No significant associations were observed between PFS/ OS and these patterns of ctDNA changes. This also applied to PFS/ OS and percent changes in ctDNA levels pre-XRT to post-XRT. These results are limited by sample size, however.
Nevertheless, the presence of specific mutations appeared to correlate with outcome. The reappearance of the driver mutations post-therapy was associated with shorter PFS. APC/ARID1A mutations present in the post-XRT blood sample correlated with shorter PFS after adjustment for tumour histology and stage. Likewise, NF1 mutations identified in post-XRT samples were associated with shorter OS after adjustment of tumour histology and stage. The final analysis of the larger cohort might be required to achieve significance for additional prognostic patterns.
REFERENCES
- Aisner DL et al., Expanded genomic testing in lung adenocarcinomas expands the survival benefit. J Clin Oncol 34, 2016 (suppl; abstr 11510)
- Zill OA et al., Somatic genomic landscape of over 15,000 patients with advanced-stage cancer from clinical next-generation sequencing analysis of circulating tumor DNA. J Clin Oncol 34, 2016 (suppl; abstr LBA11501)
- Lin SH et al., Circulating tumor DNA as a noninvasive tool to identify patients at risk for recurrence after chemoradiotherapy in stage III nonsmall cell lung cancer. J Clin Oncol 34, 2016 (suppl; abstr 8553)