Potential role of the microbiome in carcinogenesis and response to checkpoint inhibition
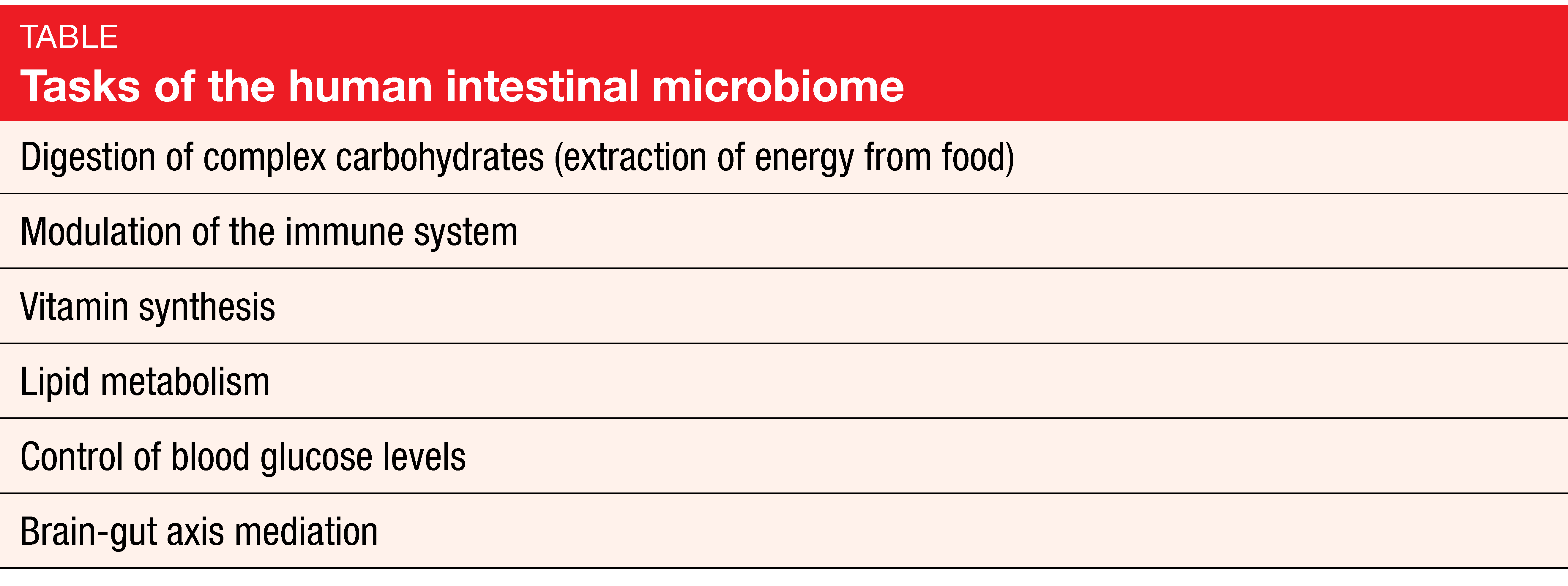
Bacteria are of eminent importance for health and the functioning of the human body that contains more bacterial than human cells, at a ratio of 1.3 to 1 [1, 2]. More than 10,000 different microbe species have been identified in the body. Ten to 100 trillion symbiotic microbial cells are harbored by each individual, primarily in the gut. Their genes outnumber the genes in the human genome by approximately 100 to 1. The intestinal microbiome fulfills a number of important tasks relating to metabolism and modulation of the immune system (Table) [3].
Here, the concept of dysbiosis comes into play, i. e. the persistent departure of the host symbiotic microbial ecosystem from the health-associated, homeostatic state towards a cancer-promoting and/or -sustaining phenotype [4, 5]. Nevertheless, dysbiosis appears to be specific to the individual, the disease, and the niches. A similar “core microbiome” was found at the phylum level (Bacteroidetes and Firmicutes), although at lower taxonomic levels, differences prevailed in apparently healthy individuals.
Microbes and cancer
Microbes can be both commensals and pathogens, with a potential role in the etiopathogenesis of cancer. Many of the most common cancers are at least partly attributable to infection. Estimates range from 20 % in lymphomas and leukemias to almost 100 % in cervical cancer [6]. Other types of cancer that are less obviously related to infections might also be triggered or promoted by dysfunctional bacterial growth. The first report suggesting the importance of microbiota in bowel carcinoma was published in 1969 [7]. Meanwhile, various studies have established a relationship between Fusobacterium nucleatum and colorectal cancer. This pathogen was shown to be enriched in colorectal cancer as compared to normal tissues [8, 9]. Distinct gut microbiome patterns correlate with consensus molecular subtypes [10], and an association was found with specific anatomic location, stage, and molecular features [11, 12]. Most recently, Fusobacterium was also detected in colorectal liver metastases by metagenomic sequencing, visualized by in situ hybridization and isolated by culture, suggesting that colorectal tumor cells provide a specific niche for this microbe [13]. These data have led to investigation of a causative role for Fusobacterium and the gut microbiome in general in CRC development and progression.
Models of underlying mechanisms
Specific microbes have been shown to modulate numerous hallmarks of cancer through diverse mechanisms [14]. All of these result in prolonged host cell survival, enhanced replicative capacity, and dedifferentiation. Two conceptual frameworks have been suggested that best describe the promotion of carcinogenesis by the human microbiome. The alpha-bug hypothesis states that certain microbiome members possessing unique virulence traits are directly pro-oncogenic and, in addition, capable of remodeling the colonic bacterial community, eventually inducing colon cancer [15]. On the other hand, according to the driver-passenger hypothesis, driver bacteria are outcompeted by passenger bacteria as mutations accumulate and adenoma turns into carcinoma [16].
In 2019, the International Cancer Microbiome Consortium published a consensus statement on the role of the human microbiome in carcinogenesis [17]. The panel concluded that, despite mechanistic and supporting evidence from animal and human studies, there is currently no direct evidence that the human commensal microbiome is a key determinant in the etiopathogenesis of cancer. A principal deciding factor in this was the lack of large longitudinal cohort studies. However, at the same time, expert opinion was that the microbiome, alongside environmental factors and an epigenetically/genetically vulnerable host, represents one apex of a tripartite, multidirectional interactome that drives carcinogenesis.
Data from large, international, longitudinal cohort studies should therefore be a future research priority to confirm the role of the human microbiome in the etiopathogenesis of cancer. There is also a need to put an increased focus on interventional studies, integration of data with other oncology research, and standardization as well as transparency in reporting microbiome research.
Antibiotics and response to immunotherapy
From the therapeutic point of view, the microbiome has emerged as a biomarker of response to immune checkpoint inhibition (ICI). As the mouse model suggests, the absence of an intact gut microbiome negatively affects the efficacy of immunotherapy [18, 19]. This implies that the use of antibiotics plays an important role here. Indeed, a recently published paper by Elkrief et al. summarized multiple clinical studies including more than 1,800 patients that demonstrated the negative predictive impact of broad-spectrum antibiotics in ICI-treated patients [20]. The data indicated that the deleterious effect is particularly pronounced if antibiotics are prescribed preceding (rather than during) immunotherapy. Thus, pre-ICI antibiotic treatment represents one risk factor of resistance through altering the diversity and composition of the intestinal microbiota. Microbiome profiling revealed that higher diversity and certain immunogenic bacteria such as Akkermansia, Firmicutes and Bifidobacterium were overrepresented in NSCLC and melanoma patients who responded to immunotherapy [18, 21, 22]. The current mechanism linking these immunogenic bacteria to CD4+ and CD8+ T cell priming appears to be involved in the reduction of ICI activity.
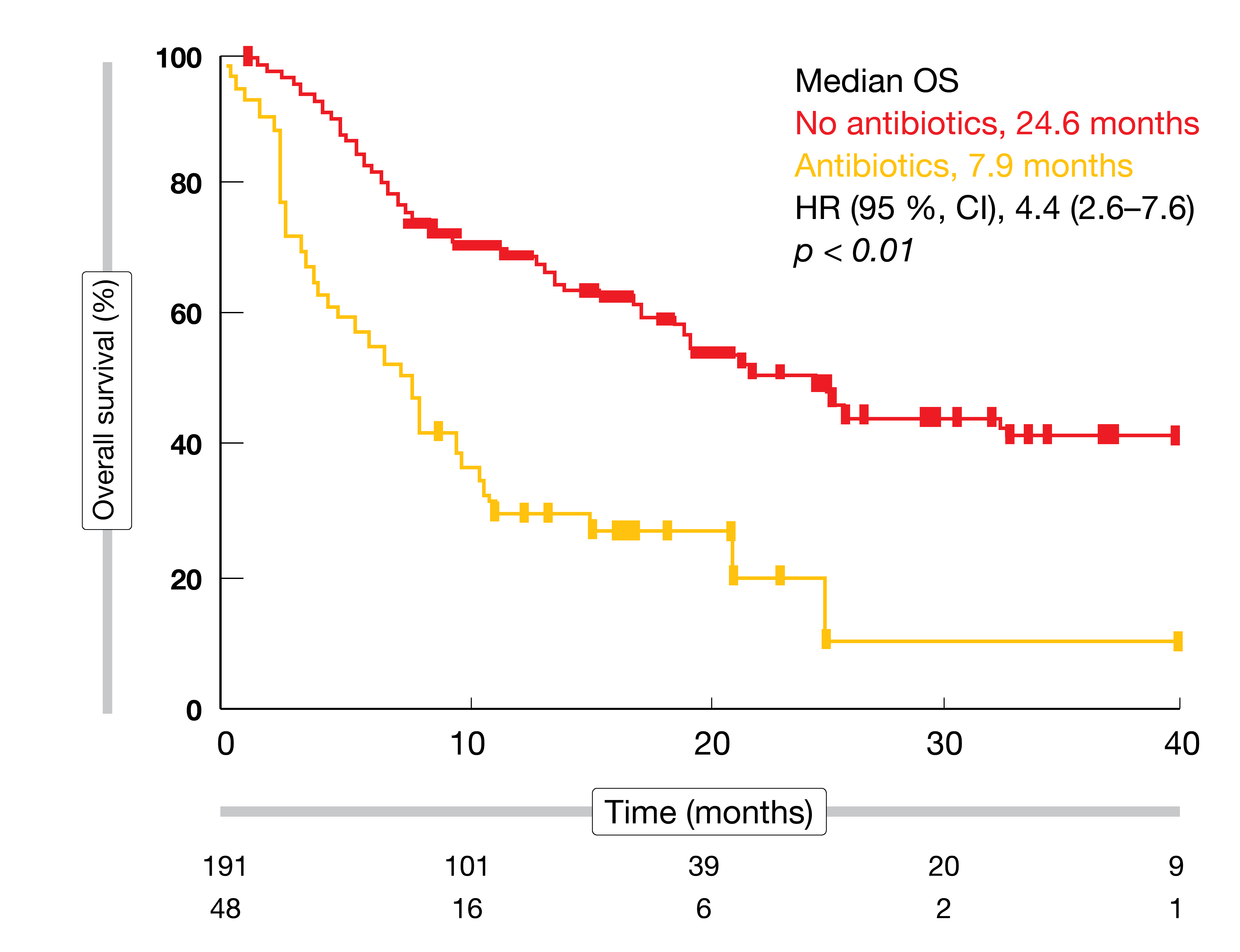
Survival outcomes can change considerably as a function of anti-microbial pretreatment. Derosa et al. showed that in patients with renal cell carcinoma, use of antibiotics compared with no use was associated with an increased risk of primary progressive disease (75 % vs. 22 %; p < 0.01), shorter PFS (1.9 vs. 7.4 months; p < 0.01], and shorter OS (17.3 vs. 30.6 months; p = 0.03) [23]. In patients with lung cancer, PFS was decreased (1.9 vs. 3.8 months; p = 0.03), as well as OS (7.9 vs. 24.6 months; p < 0.01; Figure). Controversy remains, however, as antibiotic treatment might simply constitute a surrogate marker of unfit or immunodeficient patients.
Figure: Impact of use of antibiotics on overall survival results in patients with lung cancer
How to restore microbiome health?
Nevertheless, modification of the gut microbiome is already being evaluated as an innovative therapeutic opportunity in immuno-oncology. Multiple clinical trials are ongoing that attempt to favorably shift the microbiome composition. The methods assessed here include fecal microbiota transplantation (FMT) from ICI responders or healthy donors, bacterial consortia/mixtures, high fiber/whole food dietary intervention, and co-administration of charcoal-based capsules with antibiotics. Patients with a range of tumor types including melanoma, gastric cancer and lung cancer have been enrolled. A phase I trial that assessed FMT and re-induction of anti-PD-1 therapy in patients with refractory metastatic melanoma found that FMT was safe [24]. According to the conclusion of the authors, this treatment may alter recipient gut microbiota to resemble that of a responder donor, with resulting intra-tumoral T-cell activity which translated to a clinical and radiological benefit.
Also, there may be an association between the gut microbiome and immune-mediated colitis. Wang et al. reported the first case series of ICI-associated colitis successfully treated with FMT [25]. The researchers observed reconstitution of the gut microbiome and a relative increase in the proportion of regulatory T cells within the colonic mucosa. These preliminary data provide evidence that modulation of the gut microbiome may abrogate ICI-associated colitis.
General recommendations
Elkrief et al. recommended various steps promoting the judicious use of antibiotics in ICI-treated patients [20]. If there is high suspicion of a bacterial infection in a given patient, the diagnosis should be confirmed with appropriate testing (e. g., blood cultures, imaging) before the prescription of an antibiotic. Narrow-spectrum agents should be preferred, and a shorter course should be followed if possible. Infectious disease consultations can be considered for antibiotic stewardship. The use of antibiotics should be avoided for one month preceding immunotherapy.
Overall, although the clinical significance of the gut microbiome still requires further elucidation, it constitutes a potential biomarker that needs to be included in personalized immuno-oncology trials.
Source:
Lecture “The importance of the microbiome in cancer”, Paolo Nuciforo, MD, PhD, Vall d’Hebron Institute of Oncology, Barcelona, Spain; ESMO Congress 2019, 29th September
Lecture “Working on the microbiome”, Bertrand Routy, MD, PhD, University of Montreal Medical Center, Canada; ESMO Congress 2019, 28th September
REFERENCES
- Sender R et al., Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 2016; 164(3): 337-340
- Turnbaugh PJ et al., The human microbiome project: exploring the microbial part of ourselves in a changing world. Nature 2007; 449(7164): 804-810
- Kinross JM et al., Gut microbiome-host interactions in health and disease. Genome Med 2011; 3(3): 14
- Hall AB et al., Human genetic variation and the gut microbiome in disease. Nature Rev 2017; 18(11): 690-699
- Herbst RS et al., Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 2016; 387(10027): 1540-1550
- Gilbert JA et al., Current understanding of the human microbiome. Nature Med 2018; 24(4):392-400
- www.cancer.org/canceratlas
- Aries V et al., Bacteria and the aetiology of cancer of the large bowel. Gut 1969; 10(5): 334-335
- Kostic AD et al., Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 2012; 22(2): 292-298
- Castellarin M et al., Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res 2012; 22(2): 299-306
- Purcell RV et al., Distinct gut microbiome patterns associate with consensus molecular subtypes of colorectal cancer. Scientific Reports 2017; 7(1): 11590
- Mima K et al., Fusobacterium nucleatum in colorectal carcinoma tissue according to tumor location. Clin Transl Gastroenterol 2016; 7(11): e200
- Mima et al., Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut 2016; 65(12):1973-1980
- Bullman S et al., Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 2017; 358(6369): 1443-1448
- Fulbright L et al., The microbiome and the hallmarks of cancer. PLoS Pathog 2017; 13(9): e1006480.
- Sears CL & Pardoll DM, Perspective: alpha-bugs, their microbial partners, and the link to colon cancer. J Infect Dis 2011; 203(3): 306-311
- Tjalsma H et al., A bacterial driver-passenger model for colorectal cancer: beyond the usual suspects. Nat Rev Microbiol 2012; 10(8): 575-582
- Scott AJ et al., International Cancer Microbiome Consortium consensus statement on the role of the human microbiome in carcinogenesis. Gut 2019; 68: 1624-1632
- Routy B et al., Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 2018; 359(6371): 91-97
- Vétizou M et al., Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 2015; 350(6264): 1079-84
- Elkrief A et al., The negative impact of antibiotics on outcomes in cancer patients treated with immunotherapy: a new independent prognostic factor? Ann Oncol 2019 Jul 3. pii: mdz206. doi: 10.1093/annonc/mdz206. [Epub ahead of print]
- Gopalakrishnan V et al., Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 2018; 359(6371): 97-103
- Derosa L et al., Negative association of antibiotics on clinical activity of immune checkpoint inhibitors in patients with advanced renal cell and non-small-cell lung cancer. Ann Oncol 2018; 29(6): 1437-1444
- Baruch EN et al. Fecal microbiota transplantation (FMT) and re-induction of anti-PD-1 therapy in refractory metastatic melanoma patients – preliminary results from a phase I clinical trial (NCT03353402). AACR Annual Meeting 2019, abstract CT042
- Wang Y et al., Fecal microbiota transplantation for refractory immune checkpoint inhibitor-associated colitis. Nat Med 2019; 25(1): 188