Disease monitoring using circulating cell-free tumour DNA
In Caucasian patients, mutations of the EGFR gene occur in 10 % to 15 % of adenocarcinomas of the lung. “These tumours depend on EGFR signalling for growth and survival,” explained Anna Buder, MSc, Institute of Cancer Research, Department of Medicine I, Comprehensive Cancer Center, Medical University of Vienna, Vienna, Austria. As these patients are sensitive to treatment with EGFR tyrosine kinase inhibitors (TKIs), identification of oncogenic driver mutations in adenocarcinoma has become a standard procedure in diagnostic testing.
Access to mutations via blood sampling
The EGFR gene contains 28 exons. Activating EGFR mutations are found in exons 18 to 21, which code for the tyrosine kinase domain of EGFR. “Although mutations occur throughout this domain, only some of them confer sensitivity to EGFR TKIs,” Ms. Buder noted. The most prevalent mutations are lesions in exon 19, which make up 45 % of sensitising mutations. Point mutations in exon 21 are also very common, especially L858R. Nucleotide substitutions can be identified in exon 18, and there can be in-frame insertions in exon 20. Acquired resistance caused by the T790M mutation develops in more than half of TKI-treated patients, necessitating changes in their treatment. According to an analysis conducted at the Institute of Cancer Research in Vienna, 35 % of 167 patients tested negative for the T790M mutation and also stayed negative during follow-up, while 65 % developed the T790M mutation over time.
Mutations are highly specific, as they are present in cancer cells, but not in normal body cells. Evaluation of somatic mutations is done either by conventional tumour tissue examination or by analysis of blood or body fluids; here, the testing can be performed using circulating cell-free tumour DNA, circulating tumour cells, or exosomes. Of course, liquid biopsies offer the advantage of minimal invasiveness, as they are based on normal blood sampling.
Circulating cell-free tumour DNA (ctDNA) consists of DNA fragments with a half-life of approximately 2 hours. Only traces of cell-free DNA are present in the plasma, but they are highly specific for the tumour. The length of these fragments is typically 120 – 200 base pairs, with 180 base pairs representing the length of the DNA chain within one nucleosome [1].
Measuring systems
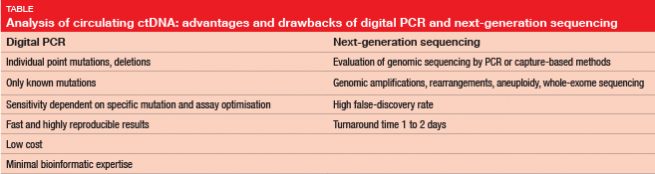
Several options are available for ctDNA testing. Digital polymerase chain reaction (PCR) is one possibility, such as droplet digital PCR (ddPCR), although this only allows for the search for known mutations (Table). “Primers that target specific mutations have to be used for this test,” Ms. Buder explained. Next-generation sequencing (NGS), on the other hand, can be used to evaluate entire genomic regions, and to detect de-novo mutations. On the downside, NGS has a long turnaround time of several days, and a high false-discovery rate. “Complicated bioinformatic techniques are required to exclude artefacts of false-positives,” Ms. Buder said.
At the Institute of Cancer Research, activating and resistance EGFR mutations are analysed from base-line blood samples and follow-up blood samples that are drawn every 1 to 3 months. “This allows for minimally invasive assessment of the response to treatment and the development of resistance.” For ddPCR, cell-free tumour DNA is extracted from the plasma and partitioned into droplets, each of which contains 0 or 1 molecule of the target DNA [2]. PCR is performed on each droplet. Droplets containing mutant and wild-type DNA emit differently coloured signals, which enables analysis and quantification.
High sensitivity of ddPCR
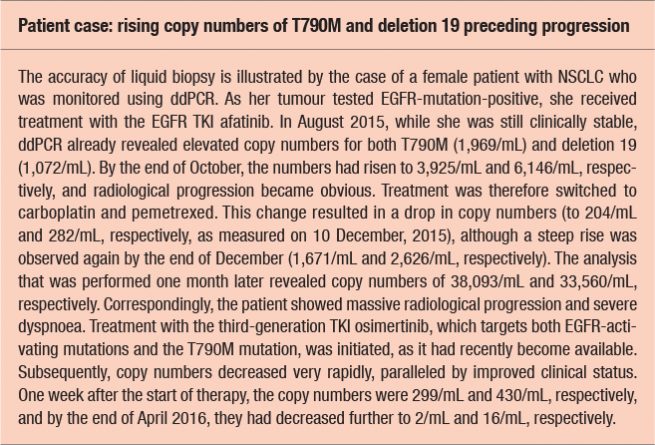
As the amount of ctDNA frequently correlates with tumour load, increasing copy numbers of activating mutations or resistance mutations might be indicative of disease progression and can be observed prior to radiological and/or clinical deterioration (see Box). Likewise, decreases suggest response to therapy. Copy numbers can range from zero to tens of thousands.
“We compared the T790M test results obtained with ddPCR with the results of the real-time-PCR-based Cobas test, which is being used by many Austrian hospitals,” Ms. Buder reported. Patients with very low numbers of DNA-mutated fragments in particular tended to be positive according to ddPCR, but negative according to Cobas. “It appears that Cobas is less sensitive than ddPCR, which detects EGFR T790M with high specificity and sensitivity,” concluded Ms. Buder. This is an important finding, because a considerable proportion of patients carries only very small amounts of mutated DNA in their plasma. For patients with 1 to 20 copies/mL, the detection rate is 37 % with ddPCR, while nearly all of these will test negative with the currently used Cobas test. Moreover, the researchers compared liquid biopsy and tissue re-biopsy with regard to the T790M mutation rate. This analysis yielded a high concordance rate of approximately 80 %.
Overall, liquid biopsy appears to be an appropriate method for identification of actionable alterations and selection of TKI therapy. Plasma ddPCR is a powerful tool for early detection of resistance mechanisms. “Liquid biopsy has the potential to replace tissue biopsy in the future,” Ms. Buder summarised.
REFERENCES
- Jahr S et al., DNA fragments in the blood plasma of cancer patients: quantitation and evidence for their origin from apoptotic and necrotic cells. Cancer Res 2001; 61(4): 1659-1665
- Oxnard GR et al., Non-invasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res 2014; 20(6): 1698-1705